Natural Gradient shares some common ideas with the high-level policy-based methods like Trust Region Policy Optimization (TRPO), Proximal Policy Optimization (PPO) and etc. The basic idea of natural policy gradient is to use the curvature information of the of the policy’s distribution over actions in the weight update. We will explain in detail how it is formulated and applied in this post.
1. Minorize-Maximization, MM
The MM algorithm is not an algorithm, but a strategy for constructing optimization algorithms.The main idea of MM (minorization-maximization) algorithms is that, intuitively, for a maximization problem, we first find a approximated lower bound of the original objective as the surrogate objective and then maximize the approximated lower bound so as to optimize the original objective. (And vice versa for minimization problems) In minimization MM stands for Majorize–Minimize, and in maximization MM stands for Minorize–Maximize. The EM algorithm can be thought as a special case of MM.
The explanation of MM (Reference:Wikipedia)
Take the minorize-maximization version for example, let ![]() be the objective concave function to be maximized. At the
be the objective concave function to be maximized. At the ![]() step of the algorithm, the constructed function
step of the algorithm, the constructed function ![]() is the surrogate function at
is the surrogate function at ![]() if
if
![]()
![]()
Now, we get
![]()
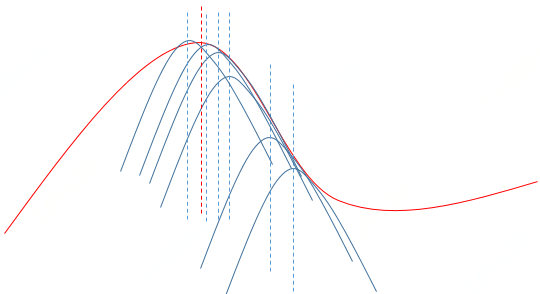 Benifit of MM that substituting a simple optimization problem for a difficult problem can turn a non-differential problem into a smooth problem and linearize an optimization problem.
Benifit of MM that substituting a simple optimization problem for a difficult problem can turn a non-differential problem into a smooth problem and linearize an optimization problem.
2. Fisher Information Matrix
Suppose we are using a model parameterized by parameter vector ![]() to model a distribution
to model a distribution ![]() , the parameter
, the parameter ![]() can be learned by maximizing the likelikhood
can be learned by maximizing the likelikhood ![]() with respect to
with respect to ![]() . To measure to the goodness of this estimation of
. To measure to the goodness of this estimation of ![]() , we define the gradient of log likelihood function of
, we define the gradient of log likelihood function of ![]() as the score function:
as the score function:
![]()
![Rendered by QuickLaTeX.com \[\begin{array}{l}\mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {s\left( \theta \right)} \right] = \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {{\nabla _\theta }\log p\left( {x\left| \theta \right.} \right)} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \int {{\nabla _\theta }\log p\left( {x\left| \theta \right.} \right)} p\left( {x\left| \theta \right.} \right)dx\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \int {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} p\left( {x\left| \theta \right.} \right)dx\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \int {{\nabla _\theta }p\left( {x\left| \theta \right.} \right)dx} \\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = {\nabla _\theta }\int {p\left( {x\left| \theta \right.} \right)dx} \\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = {\nabla _\theta }1\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = 0\end{array}\]](http://www.liaoyong.net/wp-content/ql-cache/quicklatex.com-ea3dd6e073a98204e317cab1ac5933ad_l3.png)
Here, the covariance defined as an uncertainty measure around the expected estimate is used to measure how certain we are to the estimate:
![]()
![]()
![]()
Fisher and Hessian
Feature: the negative expected Hessian of log likelihood is equal to the Fisher Information Matrix
Proof:
![Rendered by QuickLaTeX.com \[\begin{array}{l}{{\rm{H}}_{\log p\left( {x\left| \theta \right.} \right)}} = Jacobian\left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right)\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \frac{{{{\rm{H}}_{p\left( {x\left| \theta \right.} \right)}}p\left( {x\left| \theta \right.} \right) - {\nabla _\theta }p\left( {x\left| \theta \right.} \right){\nabla _\theta }p{{\left( {x\left| \theta \right.} \right)}^T}}}{{p\left( {x\left| \theta \right.} \right)p\left( {x\left| \theta \right.} \right)}}\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \frac{{{{\rm{H}}_{p\left( {x\left| \theta \right.} \right)}}p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)p\left( {x\left| \theta \right.} \right)}} - \frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right){\nabla _\theta }p{{\left( {x\left| \theta \right.} \right)}^T}}}{{p\left( {x\left| \theta \right.} \right)p\left( {x\left| \theta \right.} \right)}}\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \frac{{{{\rm{H}}_{p\left( {x\left| \theta \right.} \right)}}}}{{p\left( {x\left| \theta \right.} \right)}} - \left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right){\left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right)^T}\end{array}\]](http://www.liaoyong.net/wp-content/ql-cache/quicklatex.com-a2f680f60c5b08618f8d22466075b0d1_l3.png)
![Rendered by QuickLaTeX.com \[\begin{array}{l}\mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {{{\rm{H}}_{\log p\left( {x\left| \theta \right.} \right)}}} \right] = \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\frac{{{{\rm{H}}_{p\left( {x\left| \theta \right.} \right)}}}}{{p\left( {x\left| \theta \right.} \right)}} - \left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right){{\left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right)}^T}} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\frac{{{{\rm{H}}_{p\left( {x\left| \theta \right.} \right)}}}}{{p\left( {x\left| \theta \right.} \right)}}} \right] - \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right){{\left( {\frac{{{\nabla _\theta }p\left( {x\left| \theta \right.} \right)}}{{p\left( {x\left| \theta \right.} \right)}}} \right)}^T}} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \int {\frac{{{{\rm{H}}_{p\left( {x\left| \theta \right.} \right)}}}}{{p\left( {x\left| \theta \right.} \right)}}} p\left( {x\left| \theta \right.} \right)dx - \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {{\nabla _\theta }\log p\left( {x\left| \theta \right.} \right){\nabla _\theta }\log p{{\left( {x\left| \theta \right.} \right)}^{\rm{T}}}} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = {{\rm{H}}_{\int {p\left( {x\left| \theta \right.} \right)dx} }} - \mathbb{F}\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = {{\rm{H}}_1} - \mathbb{F}\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = - \mathbb{F}\end{array}\]](http://www.liaoyong.net/wp-content/ql-cache/quicklatex.com-4775d8cadfc61a0a7d5743cb0135e1b2_l3.png)
![]()
3. Fisher Information Matrix and KL-divergence
KL divergence maps to our notion of what a probability distance should look like: it measures directly in terms of how probability density functions are defined, that is, differences in density value at a bunch of points over which the distribution is defined.
The relationship between Fisher Information Matrix and KL-divergence is that FIM defines the local curvature in distribution space for which KL-divergence is the metric. That is, FIM F is the Hessian of KL-divergence between two distributions ![]() and
and ![]() , with respect to
, with respect to ![]() , evaluated at
, evaluated at ![]() .
.
This can be proved as follow:
![Rendered by QuickLaTeX.com \[\begin{array}{l}{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta '} \right.} \right)} \right.} \right] = \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\log p\left( {x\left| \theta \right.} \right)} \right] - \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\log p\left( {x\left| {\theta '} \right.} \right)} \right]\\{\nabla _{\theta '}}{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta '} \right.} \right)} \right.} \right] = {\nabla _{\theta '}}\mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\log p\left( {x\left| \theta \right.} \right)} \right] - {\nabla _{\theta '}}\mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {\log p\left( {x\left| {\theta '} \right.} \right)} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = - \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {{\nabla _{\theta '}}\log p\left( {x\left| {\theta '} \right.} \right)} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = - \int {p\left( {x\left| \theta \right.} \right)} {\nabla _{\theta '}}\log p\left( {x\left| {\theta '} \right.} \right)dx\\\nabla _{\theta '}^2{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta '} \right.} \right)} \right.} \right] = - \int {p\left( {x\left| \theta \right.} \right)} \nabla _{\theta '}^2\log p\left( {x\left| {\theta '} \right.} \right)dx\\{\rm{H}_{{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta '} \right.} \right)} \right.} \right]}} = - \int {p\left( {x\left| \theta \right.} \right)} \nabla _{\theta '}^2\log p\left( {x\left| {\theta '} \right.} \right)\left| {_{\theta ' = \theta }} \right.dx\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = - \int {p\left( {x\left| \theta \right.} \right)} {\rm{H}_{\log p\left( {x\left| \theta \right.} \right)}}dx\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = - \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} \left[ {{\rm{H}_{\log p\left( {x\left| \theta \right.} \right)}}} \right]\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = \mathbb{F}\end{array}\]](http://www.liaoyong.net/wp-content/ql-cache/quicklatex.com-56e2c6e7204fb3fb20248e59d212f464_l3.png)
Using Taylor expansion, when
![Rendered by QuickLaTeX.com \[\begin{array}{l}{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta + d} \right.} \right)} \right.} \right] \approx {\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| \theta \right.} \right)} \right.} \right] + {\left( {{\nabla _{\theta '}}{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta '} \right.} \right)} \right.} \right]\left| {_{\theta ' = \theta }} \right.} \right)^{\rm{T}}}d + \frac{1}{2}{d^{\rm{T}}}\rm{F}d\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} = {\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| \theta \right.} \right)} \right.} \right] - \mathop \mathbb{E}\limits_{p\left( {x\left| \theta \right.} \right)} {\left[ {{\nabla _\theta }\log p\left( {x\left| \theta \right.} \right)} \right]^{\rm{T}}} + \frac{1}{2}{d^{\rm{T}}}\rm{F}d\end{array}\]](http://www.liaoyong.net/wp-content/ql-cache/quicklatex.com-f10661fd8cd0e08f1442e0e78da4f7d8_l3.png)
![]()
![]()
![]()
To solve this minimization problem with constraint, we convert it to unconstrained problem and rewrite the equation as
![Rendered by QuickLaTeX.com \[\begin{array}{l}{d^*} = \mathop {\arg \min }\limits_d \mathcal{L}\left( {\theta + d} \right) + \lambda \left( {{\rm{KL}}\left[ {p\left( {x\left| \theta \right.} \right)\left\| {p\left( {x\left| {\theta + d} \right.} \right)} \right.} \right] - c} \right)\\{\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} {\kern 1pt} \approx \mathop {\arg \min }\limits_d \mathcal{L}\left( \theta \right) + {\nabla _\theta }\mathcal{L}{\left( \theta \right)^{\rm{T}}}d + \frac{1}{2}\lambda {d^{\rm{T}}}{\rm{F}}d - \lambda c\end{array}\]](http://www.liaoyong.net/wp-content/ql-cache/quicklatex.com-eda8f51a69642df462b5ff4054bafb85_l3.png)
![]()
![]()
4. Natural Policy Gradient
The Natural Gradient is defined as
![]()
Natural Gradient provides information about curvature and a way to directly control movement of your model in predicted distribution space, as separate from movement of your model in loss space
More explanations of Natural Gradient (reference…).
Standard Gradient Decent:
Let’s say we have a neural network, parameterized by some vector of parameters. We want to adjust the parameters of this network, so the output of our network changes in some way. In most cases, we will have a loss function that tells our network how it’s output should change.
Using backpropagation, we calculate the derivatives of each parameter with respect to our loss function. These derivatives represent the direction in which we can update our parameters to get the biggest change in our loss function, and is known as the gradient. We can then adjust the parameters in the direction of the gradient, by a small distance, in order to train our network.
A slight problem appears in how we define “a small distance”.
|
1 2 |
#distance with two parameters (a,b) distance = sqrt(a^2 + b^2) |
However, defining “distance” in terms of how much our parameters is adjusted isn’t always correct. To visualize this, let’s take a look at two pairs of Gaussian distributions.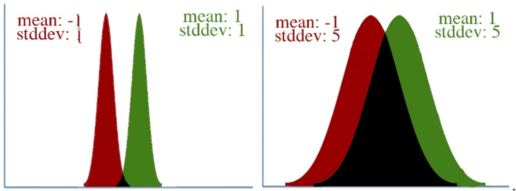 In both distributions, the mean is changed from -1 to 1, a distance of two. However, it’s clear that the first distribution changed far more than the second.This leads to a key insight: our gradient measures how much our output is affected by changing a parameter. However, this affect on the output must be seen in context: a shift of +2 in the first distribution means a lot more than a shift of +2 in the second one.
In both distributions, the mean is changed from -1 to 1, a distance of two. However, it’s clear that the first distribution changed far more than the second.This leads to a key insight: our gradient measures how much our output is affected by changing a parameter. However, this affect on the output must be seen in context: a shift of +2 in the first distribution means a lot more than a shift of +2 in the second one.
What the natural gradient does is redefine the “small distance” we update our parameters by. Not all parameters are equal. Rather than treating a change in every parameter equally, we need to scale each parameter’s change according to how much it affects our network’s entire output distribution.
How do we do that?
First off, let’s define a new form of distance, that corresponds to distance based on KL divergence, a measure of how much our new distribution differs from our old one. We do this by defining a metric matrix, which allows us to calculate the distance of a vector according to some custom measurement.
For a network with 5 parameters, our metric matrix is 5×5. To compute the distance of a change in parameters delta using metric, we use the following:
|
1 2 3 4 |
total_distance = 0 for i in xrange(5): for j in xrange(5): total_distance += delta[i] * delta[i] * metric[i][j] |
If our metric matrix is the identity matrix, the distance is the same as if we just used Euclidean distance.However, most of the time our metric won’t be the identity matrix. Having a metric allows our measurement of distance to account for relationships between the various parameters. As it turns out, we can use the Fisher Information Matrix as a metric, and it will measure the distance of delta in terms of KL divergence.The Fisher Information Matrix is the second derivative of the KL divergence of our network with itself. For more information on why, see this article.
Now we’ve got a metric matrix that measures distance according to KL divergence when given a change in parameters.
With that, we can calculate how our standard gradient should be scaled.
|
1 |
natural_grad = inverse(fisher) * standard_grad |
Notice that if the Fisher is the identity matrix, then the standard gradient is equivalent to the natural gradient. This makes sense: using the identity as a metric results in Euclidean distance, which is what we were using in standard gradient descent.
Finally, I want to note that computing the natural gradient may be computationally intensive. In an implementation of TRPO, the conjugate gradient algorithm is used to solve for natural_grad in “natural_grad * fisher = standard_grad”.
Pseudocode
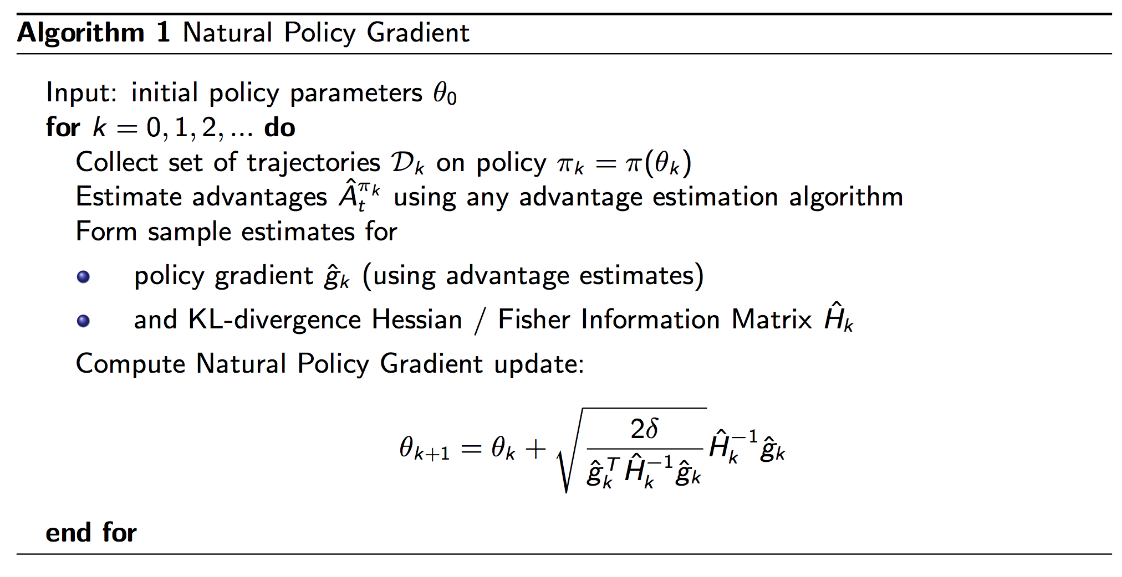 Implementation
Implementation
|
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 |
import tensorflow.compat.v1 as tf tf.disable_v2_behavior() # to make it possible to run 1.X code in tf2 import numpy as np import gym import random def policy_gradient(): with tf.variable_scope("policy"): params = tf.get_variable("policy_parameters", [4, 2]) state = tf.placeholder("float", [None, 4], name="state") # NOTE: have to specify shape of actions so we can call # get_shape when calculating g_log_prob below actions = tf.placeholder("float", [200, 2], name="actions") advantages = tf.placeholder("float", [None,], name="advantages") linear = tf.matmul(state, params) #shape: [None, 2] probabilities = tf.nn.softmax(linear) #shape: [None, 2] my_variables = tf.trainable_variables() # calculate the probability of the chosen action given the state # shape: [200, 2] ---> [1, 2] the probs of two actions action_log_prob = tf.log(tf.reduce_sum( tf.multiply(probabilities, actions), reduction_indices=[1])) # calculate the gradient of the log probability at each point in time # NOTE: doing this because tf.gradients only returns a summed version action_log_prob_flat = tf.reshape(action_log_prob, (-1,)) #shape:(2,) g_log_prob = tf.stack( [tf.gradients(action_log_prob_flat[i], my_variables)[0] for i in range(action_log_prob_flat.get_shape()[0])]) g_log_prob = tf.reshape(g_log_prob, (200, 8, 1)) # calculate the policy gradient by multiplying by the advantage function # shape:(200, 8, 1) g = tf.multiply(g_log_prob, tf.reshape(advantages, (200, 1, 1))) # sum over time g = 1.00 / 200.00 * tf.reduce_sum(g, reduction_indices=[0]) #shape:(8, 1) F2 = tf.map_fn(lambda x: tf.matmul(x, tf.transpose(x)), g_log_prob) F = 1.0 / 200.00 * tf.reduce_sum(F2, reduction_indices=[0]) #T=200 # calculate inverse of positive definite clipped F # NOTE: have noticed small eigenvalues (1e-10) that are negative, # using SVD to clip those out, assuming they're rounding errors S, U, V = tf.svd(F) atol = tf.reduce_max(S) * 1e-6 S_inv = tf.divide(1.0, S) S_inv = tf.where(S < atol, tf.zeros_like(S), S_inv) S_inv = tf.diag(S_inv) F_inv = tf.matmul(S_inv, tf.transpose(U)) F_inv = tf.matmul(V, F_inv) # calculate natural policy gradient ascent update F_inv_g = tf.matmul(F_inv, g) # calculate a learning rate normalized such that a constant change # in the output control policy is achieved each update, preventing # any parameter changes that hugely change the output learning_rate = tf.sqrt(tf.divide(0.001, tf.matmul(tf.transpose(g), F_inv_g))) update = tf.multiply(learning_rate, F_inv_g) update = tf.reshape(update, (4, 2)) # update trainable parameters # NOTE: whenever my_variables is fetched they're also updated my_variables[0] = tf.assign_add(my_variables[0], update) return probabilities, state, actions, advantages, my_variables def value_gradient(): with tf.variable_scope("value"): state = tf.placeholder("float", [None, 4]) new_vals = tf.placeholder("float", [None, 1]) w1 = tf.get_variable("w1", [4, 10]) b1 = tf.get_variable("b1", [10]) h1 = tf.nn.relu(tf.matmul(state, w1) + b1) w2 = tf.get_variable("w2", [10, 1]) b2 = tf.get_variable("b2", [1]) calculated = tf.matmul(h1, w2) + b2 # minimize the difference between predicted and actual diffs = calculated - new_vals loss = tf.nn.l2_loss(diffs) optimizer = tf.train.AdamOptimizer(0.1).minimize(loss) return calculated, state, new_vals, optimizer, loss def run_episode(env, policy_grad, value_grad, sess): # unpack the policy network (generates control policy) (pl_calculated, pl_state, pl_actions, pl_advantages, pl_optimizer) = policy_grad # unpack the value network (estimates expected reward) (vl_calculated, vl_state, vl_new_vals, vl_optimizer, vl_loss) = value_grad # set up the environment observation = env.reset() #shape:(4,) episode_reward = 0 total_rewards = [] states = [] actions =[] advantages = [] transitions = [] update_vals = [] n_episodes = 0 n_timesteps = 200 for t in range(n_timesteps): # calculate policy obs_vector = np.expand_dims(observation, axis=0) #shape:(1, 4) probs = sess.run(pl_calculated, feed_dict={pl_state: obs_vector}) # stochastically generate action using the policy output action = 0 if random.uniform(0, 1) < probs[0][0] else 1 # record the transition states.append(observation) action_blank = np.zeros(2) action_blank[action] = 1 actions.append(action_blank) # take the action in the environment old_observation = observation observation, reward, done, info = env.step(action) transitions.append((old_observation, action, reward)) episode_reward += reward # if the pole falls or time is up if done or t == n_timesteps - 1: for ii, trans in enumerate(transitions): obs, action, reward = trans # calculate discounted monte-carlo return future_reward = 0 future_transitions = len(transitions) - ii decrease = 1 for jj in range(future_transitions): future_reward += transitions[jj + ii][2] * decrease decrease = decrease * 0.97 obs_vector = np.expand_dims(obs, axis=0) # compare the calculated expected reward to the average # expected reward, as estimated by the value network current_val = sess.run(vl_calculated, feed_dict={ vl_state: obs_vector})[0][0] # advantage: how much better was this action than normal advantages.append(future_reward - current_val) # update the value function towards new return update_vals.append(future_reward) n_episodes += 1 # reset variables for next episode in batch total_rewards.append(episode_reward) episode_reward = 0.0 transitions = [] if done: # if the pole fell, reset environment observation = env.reset() else: # if out of time, close environment env.close() print("total_rewards: ", total_rewards) # update value function update_vals_vector = np.expand_dims(update_vals, axis=1) sess.run(vl_optimizer, feed_dict={vl_state: states, vl_new_vals: update_vals_vector}) # update control policy sess.run(pl_optimizer, feed_dict={ pl_state: states, pl_advantages: advantages, pl_actions: actions}) return total_rewards, n_episodes # generate the networks policy_grad = policy_gradient() value_grad = value_gradient() # run the training from scratch 10 times, record results for ii in range(10): #ran 10 sessions of 300 batches env = gym.make("CartPole-v0") # env = gym.wrappers.Monitor(env=env, directory="./", force=True, video_callable=False) sess = tf.InteractiveSession() sess.run(tf.global_variables_initializer()) max_rewards = [] total_episodes = [] # each batch is 200 time steps worth of episodes n_training_batches = 300 times = [] for i in range(n_training_batches): if i % 100 == 0: print(i) reward, n_episodes = run_episode(env, policy_grad, value_grad, sess) max_rewards.append(np.max(reward)) total_episodes.append(n_episodes) sess.close() |
Results
5. Summary
I will end the post with a summary graph as bellow to connection between different parts of the Natural Gradient as bellow